B2: Characterization and integrative bioinformatic modeling of metabolic and micromilieu factors promoting survival or control of Leishmania parasites
B2A (Prof. Bogdan): Metabolic and micromilieu factors regulating control and survival of Leishmania parasites
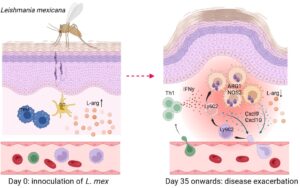
We have recently observed both in the Leishmania (L.) major and L. mexicana model that hyperexpression of arginase 1 (Arg1) accounts for a non-healing course of cutaneous leishmaniasis. Myeloid cells co-expressing Arg1 and type nitric oxide synthase (NOS2) form a replicative niche for Leishmania parasites. The availability of L-arginine at the skin site of infection turned out to be a critical factor for the development of a protective anti-Leishmania immune response and the healing of the skin lesions. Future studies will address the molecular impact of L-arginine on the transcriptome, metabolome and function of different immune cells and mesenchymal cells as well as on the skin microbiota during cutaneous leishmaniasis.
B2B (PD Dr. Kunz):
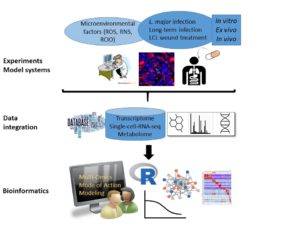
The bioinformatical part of the project aims to understand the positive and negative regulation of the anti-Leishmania immune response by metabolic and micromilieu factors using an integrative in silico modeling approach that is based on published and novel experimental data. The respective doctoral candidate will carry out parasite/immune cell interactome mapping and analyze the impact on host cell function by network analysis. A comprehensive in silico model of immune cell signaling and metabolic events in the host cell will be established that also takes into account micromilieu factors in the host cell niche during Leishmania infection. Boolean regulatory and metabolic modeling interpolates between system states in host cells, which allows to systematically simulate parasite persistence under different oxidative (e.g., presence/absence of ROS or NO), metabolic (e.g., glycolysis vs. oxidative phosphorylation) and host cell conditions (e.g., myeloid cells, fibroblasts) and at different time-points of infection. The studies will be complemented by an in silico screening of potential drug acting mechanisms of sodium chlorosum.
+Supervisors
Prof. Dr. Christian Bogdan
91054 Erlangen
- Phone number: +49 9131 85-32561
- Email: christian.bogdan@uk-erlangen.de
- Website: http://www.mikrobiologie.uk-erlangen.de
PD Dr. Meik Kunz
91058 Erlangen-Tennenlohe
- Phone number: +49 9131 85-67784
- Email: meik.kunz@fau.de
- Website: https://www.imi.med.fau.de/team/dr-meik-kunz/